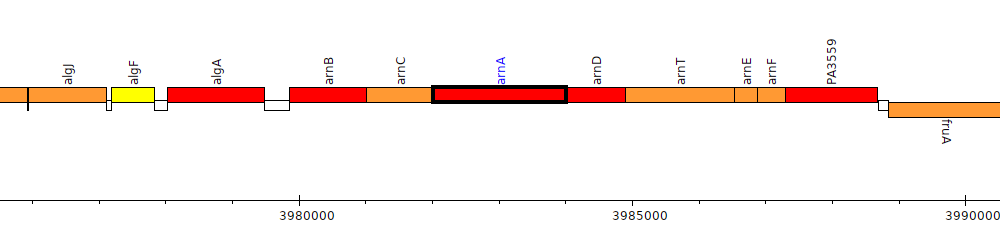
Pseudomonas aeruginosa PAO1, PA3554 (arnA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:1901760 | beta-L-Ara4N-lipid A biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
15695810 | Reviewed by curator |
| Molecular Function | GO:0016831 | carboxy-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd05257
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016742 | hydroxymethyl-, formyl- and related transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01166
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02911
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF50486
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01503 | Cationic antimicrobial peptide (CAMP) resistance | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00520 | Amino sugar and nucleotide sugar metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 319 | 652 | 8.2E-71 |
| SUPERFAMILY | SSF50486 | FMT C-terminal domain-like | IPR011034 | Formyl transferase-like, C-terminal domain superfamily | 207 | 300 | 6.75E-21 |
| Hamap | MF_01166 | Bifunctional polymyxin resistance protein ArnA [arnA]. | IPR021168 | Bifunctional polymyxin resistance protein, ArnA | 3 | 661 | 61.311974 |
| Pfam | PF00551 | Formyl transferase | IPR002376 | Formyl transferase, N-terminal | 28 | 175 | 5.8E-26 |
| CDD | cd05257 | Arna_like_SDR_e | IPR045869 | Bifunctional polymyxin resistance protein ArnA-like, extended (e) SDRs | 319 | 650 | 2.15337E-117 |
| Pfam | PF01370 | NAD dependent epimerase/dehydratase family | IPR001509 | NAD-dependent epimerase/dehydratase | 320 | 567 | 1.0E-43 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 316 | 659 | 4.8E-120 |
| PANTHER | PTHR43245 | BIFUNCTIONAL POLYMYXIN RESISTANCE PROTEIN ARNA | - | - | 319 | 651 | 1.2E-84 |
| PIRSF | PIRSF036506 | MtRNA_frt_NDPSE | IPR021168 | Bifunctional polymyxin resistance protein, ArnA | 3 | 661 | 0.0 |
| CDD | cd08702 | Arna_FMT_C | - | - | 206 | 295 | 1.81339E-32 |
| SUPERFAMILY | SSF53328 | Formyltransferase | IPR036477 | Formyl transferase, N-terminal domain superfamily | 5 | 204 | 1.83E-54 |
| FunFam | G3DSA:3.40.50.720:FF:000197 | Bifunctional polymyxin resistance protein ArnA | - | - | 316 | 660 | 0.0 |
| Pfam | PF02911 | Formyl transferase, C-terminal domain | IPR005793 | Formyl transferase, C-terminal | 204 | 280 | 4.9E-16 |
| Gene3D | G3DSA:3.40.50.12230 | - | - | - | 4 | 295 | 2.4E-82 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.