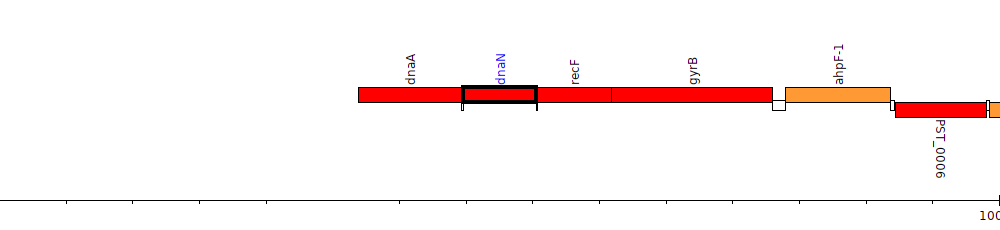
Pseudomonas stutzeri A1501, PST_0002 (dnaN)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003887 | DNA-directed DNA polymerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02768
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02768
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008408 | 3'-5' exonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02768
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:PF02768
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009360 | DNA polymerase III complex |
Inferred from Sequence Model
Term mapped from: InterPro:PF02768
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psa00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psa03430 | Mismatch repair | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psa03440 | Homologous recombination | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psa00240 | Pyrimidine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psa03030 | DNA replication | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psa01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.10.150.10 | DNA Polymerase III, subunit A, domain 2 | - | - | 1 | 123 | 1.8E-49 |
| FunFam | G3DSA:3.10.150.10:FF:000001 | Beta sliding clamp | - | - | 1 | 123 | 1.6E-53 |
| NCBIfam | TIGR00663 | JCVI: DNA polymerase III subunit beta | IPR001001 | DNA polymerase III, beta sliding clamp | 1 | 367 | 2.5E-110 |
| Gene3D | G3DSA:3.10.150.10 | DNA Polymerase III, subunit A, domain 2 | - | - | 249 | 367 | 2.1E-32 |
| PANTHER | PTHR30478 | DNA POLYMERASE III SUBUNIT BETA | IPR001001 | DNA polymerase III, beta sliding clamp | 1 | 367 | 1.3E-128 |
| SUPERFAMILY | SSF55979 | DNA clamp | IPR046938 | DNA clamp superfamily | 246 | 367 | 4.71E-37 |
| SMART | SM00480 | pol35 | IPR001001 | DNA polymerase III, beta sliding clamp | 17 | 363 | 0.0 |
| Pfam | PF02768 | DNA polymerase III beta subunit, C-terminal domain | IPR022635 | DNA polymerase III, beta sliding clamp, C-terminal | 247 | 366 | 2.5E-40 |
| CDD | cd00140 | beta_clamp | IPR001001 | DNA polymerase III, beta sliding clamp | 1 | 366 | 0.0 |
| Pfam | PF00712 | DNA polymerase III beta subunit, N-terminal domain | IPR022634 | DNA polymerase III, beta sliding clamp, N-terminal | 1 | 119 | 2.0E-41 |
| Pfam | PF02767 | DNA polymerase III beta subunit, central domain | IPR022637 | DNA polymerase III, beta sliding clamp, central | 130 | 244 | 5.9E-39 |
| Gene3D | G3DSA:3.10.150.10 | DNA Polymerase III, subunit A, domain 2 | - | - | 125 | 248 | 4.3E-40 |
| PIRSF | PIRSF000804 | DNA_pol_III_b | IPR001001 | DNA polymerase III, beta sliding clamp | 1 | 367 | 5.0E-123 |
| SUPERFAMILY | SSF55979 | DNA clamp | IPR046938 | DNA clamp superfamily | 1 | 120 | 8.59E-31 |
| SUPERFAMILY | SSF55979 | DNA clamp | IPR046938 | DNA clamp superfamily | 126 | 245 | 5.07E-30 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.