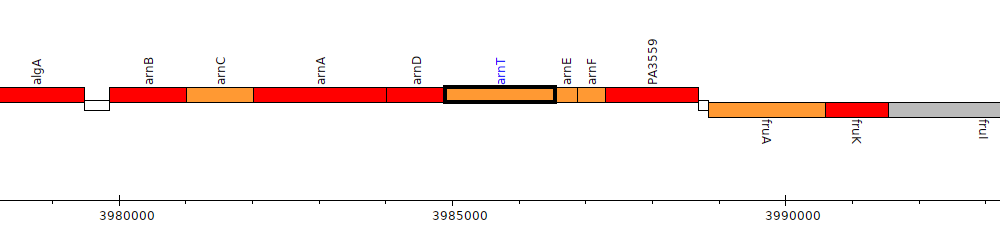
Pseudomonas aeruginosa PAO1, PA3556 (arnT)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016757 | transferase activity, transferring glycosyl groups |
Inferred from Sequence or Structural Similarity
Term mapped from: CAZy:GT83
|
ECO:0000250 sequence similarity evidence used in manual assertion |
18838391 | Reviewed by curator |
| Biological Process | GO:0046677 | response to antibiotic | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
11535604 | Reviewed by curator |
| Cellular Component | GO:0009276 | Gram-negative-bacterium-type cell wall | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
11535604 | Reviewed by curator |
| Cellular Component | GO:0016020 | membrane | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
11535604 | Reviewed by curator |
| Biological Process | GO:1901760 | beta-L-Ara4N-lipid A biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
11535604 | Reviewed by curator |
| Molecular Function | GO:0000030 | mannosyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02366
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016763 | transferase activity, transferring pentosyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01165
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006493 | protein O-linked glycosylation |
Inferred from Sequence Model
Term mapped from: InterPro:PF02366
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF02366
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01503 | Cationic antimicrobial peptide (CAMP) resistance | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_01165 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase [arnT]. | IPR022839 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase | 1 | 547 | 41.391724 |
| Pfam | PF02366 | Dolichyl-phosphate-mannose-protein mannosyltransferase | IPR003342 | Glycosyl transferase family 39/83 | 10 | 236 | 4.0E-30 |
| PANTHER | PTHR33908 | MANNOSYLTRANSFERASE YKCB-RELATED | - | - | 6 | 530 | 1.5E-81 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.