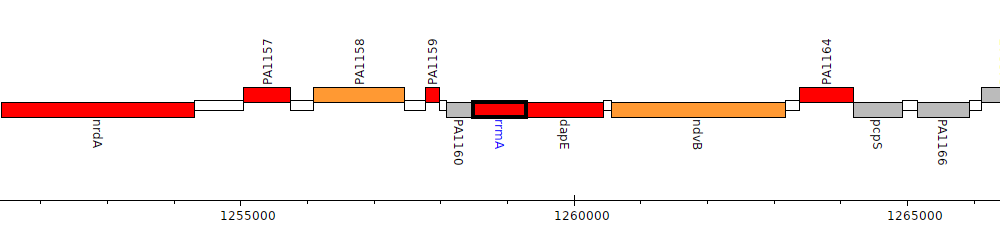
Pseudomonas aeruginosa PAO1, PA1161 (rrmA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0016070 | RNA metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0008168 | methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF018249
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53335
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF018249 | MyrA | IPR016718 | rRNA (guanine-N1-)-methyltransferase A, predicted | 1 | 267 | 1.8E-82 |
| PANTHER | PTHR43460 | METHYLTRANSFERASE | - | - | 81 | 239 | 5.4E-9 |
| Gene3D | G3DSA:3.40.50.150 | Vaccinia Virus protein VP39 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 1 | 266 | 5.4E-68 |
| SUPERFAMILY | SSF53335 | S-adenosyl-L-methionine-dependent methyltransferases | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 1 | 249 | 5.16E-39 |
| Pfam | PF13649 | Methyltransferase domain | IPR041698 | Methyltransferase domain 25 | 86 | 168 | 3.3E-10 |
| CDD | cd02440 | AdoMet_MTases | - | - | 84 | 168 | 3.18582E-6 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.