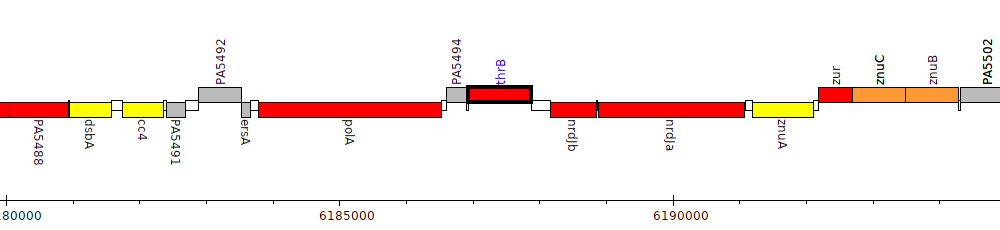
Pseudomonas aeruginosa PAO1, PA5495 (thrB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004413 | homoserine kinase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
10220164 | Reviewed by curator |
| Biological Process | GO:0009088 | threonine biosynthetic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
10220164 | Reviewed by curator |
| Molecular Function | GO:0004413 | homoserine kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00938
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009088 | threonine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00938
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Glycine, serine and threonine metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | P4-PWY | superpathway of L-lysine, L-threonine and L-methionine biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | HOMOSER-THRESYN-PWY | L-threonine biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | THRESYN-PWY | superpathway of L-threonine biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00260 | Glycine, serine and threonine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF56112 | Protein kinase-like (PK-like) | IPR011009 | Protein kinase-like domain superfamily | 5 | 306 | 1.75E-67 |
| NCBIfam | TIGR00938 | JCVI: homoserine kinase | IPR005280 | Homoserine kinase, type II | 1 | 303 | 6.0E-85 |
| Hamap | MF_00301 | Homoserine kinase [thrB]. | IPR005280 | Homoserine kinase, type II | 1 | 316 | 146.353165 |
| CDD | cd05153 | HomoserineK_II | IPR005280 | Homoserine kinase, type II | 9 | 305 | 1.54306E-102 |
| Pfam | PF01636 | Phosphotransferase enzyme family | IPR002575 | Aminoglycoside phosphotransferase | 28 | 255 | 3.4E-36 |
| PANTHER | PTHR21064 | UNCHARACTERIZED | - | - | 19 | 285 | 3.3E-24 |
| Gene3D | G3DSA:3.30.200.20 | Phosphorylase Kinase; domain 1 | - | - | 1 | 101 | 5.6E-30 |
| Gene3D | G3DSA:3.90.1200.10 | - | - | - | 105 | 314 | 6.2E-66 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.