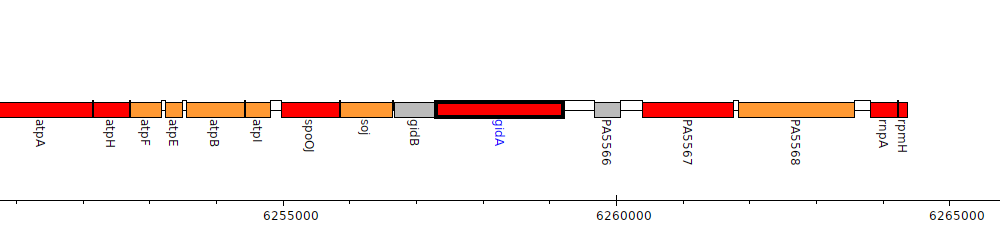
Pseudomonas aeruginosa PAO1, PA5565 (gidA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0002098 | tRNA wobble uridine modification |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P0A6U3
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17062623 | Reviewed by curator |
| Biological Process | GO:0030488 | tRNA methylation |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P0A6U3
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17062623 | Reviewed by curator |
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P0A6U3
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17062623 | Reviewed by curator |
| Biological Process | GO:0008033 | tRNA processing |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11806
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11806
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0002098 | tRNA wobble uridine modification |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00129
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 294 | 458 | 4.0E-57 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 148 | 166 | 8.4E-5 |
| Gene3D | G3DSA:1.10.150.570 | - | IPR044920 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme, C-terminal subdomain superfamily | 567 | 619 | 7.6E-24 |
| NCBIfam | TIGR00136 | JCVI: tRNA uridine-5-carboxymethylaminomethyl(34) synthesis enzyme MnmG | IPR004416 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme MnmG | 7 | 620 | 0.0 |
| FunFam | G3DSA:1.10.150.570:FF:000001 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme MnmG | - | - | 567 | 620 | 4.4E-23 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 2 | 232 | 9.5E-88 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 9 | 28 | 8.4E-5 |
| FunFam | G3DSA:3.50.50.60:FF:000002 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme MnmG | - | - | 295 | 458 | 3.1E-86 |
| PANTHER | PTHR11806 | GLUCOSE INHIBITED DIVISION PROTEIN A | IPR002218 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme MnmG-related | 6 | 625 | 0.0 |
| SMART | SM01228 | GIDA_assoc_3_2 | IPR047001 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme, C-terminal subdomain | 544 | 615 | 1.5E-41 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 7 | 261 | 1.43E-52 |
| Pfam | PF01134 | Glucose inhibited division protein A | IPR040131 | MnmG, N-terminal domain | 8 | 399 | 0.0 |
| Hamap | MF_00129 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme MnmG [mnmG]. | IPR004416 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme MnmG | 5 | 622 | 42.546913 |
| FunFam | G3DSA:3.50.50.60:FF:000010 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme MnmG | - | - | 2 | 233 | 3.2E-128 |
| Gene3D | G3DSA:1.10.10.1800 | - | - | - | 459 | 557 | 2.2E-26 |
| FunFam | G3DSA:1.10.10.1800:FF:000001 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme MnmG | - | - | 459 | 548 | 4.2E-26 |
| Pfam | PF13932 | tRNA modifying enzyme MnmG/GidA C-terminal domain | IPR026904 | tRNA uridine 5-carboxymethylaminomethyl modification enzyme MnmG, C-terminal | 402 | 614 | 4.6E-77 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.